Bayesian-Optimized HDBSCAN Clustering
Abstract
This project comprises of a clustering algorithm to identify and gate on reactions of interest measured with the Super-Enge Split-Pole Spectrograph (SE-SPS). The algorithm uses scikit’s HDBSCAN clustering function to identify the number of clusters in particle-identification (PID) plots and to prescribe each data point to a cluster or as noise. Bayesian optimization of HDBSCAN parameters was performed using scikit’s gp_minimize function, which uses Guassian Processes to increase evaluation efficiency. The algorithm was tested on experimental data with six SE-SPS magnetic field settings, with each test taking about 10 seconds to run and producing satisfactory identification of particle groups and gates around said groups. Limits to the algorithm arise if the dataset is too sparse or if there are less than three clusters, as seen in the misidentification of clusters in the 5.99 kG and 10 kG datasets.
Problem Statement
When performing experiments, many different reactions occur and it is the experiment's job to provide an apparatus to gate on the specific reaction of interest. The Super-Enge Split-Pole Spectrograph (SE-SPS) at the John D. Fox Laboratory at Florida State University uses a Particle Identification (PID) plot in order to differentiate the output particles of the reactions. The PID plot displays distinct clusters that correspond to different ejected particles: protons, deuterons, alphas, and tritons. The experimenter then needs to manually click and gate around the cluster of interest. This may seem easy, but each cluster changes size and shape depending on the angle of the detector, the magnetic field setting of the SE-SPS, as well as the beam energy. Therefore, this procedure becomes quite time consuming, since these experiment settings need to be continually tuned, requiring new gates to be made. This clustering algorithm attempts to streamline the process of identifying each particle group, omitting outliers, and performing a cut around the desired group no matter what settings are applied in the experiment. HDBSCAN was chosen for it robustness as it does not assume that the density is globally homogeneous. Instead, it explores all possible density scales which is important since clusters in the PID plot can, and often do, have different densities as reaction cross-sections (probabilities) vary. While HDBSCAN is a powerful clustering algorithm, its performance heavily depends on hyperparameter selection. Traditional grid search or random search methods are computationally expensive and may not find optimal parameters, therefore I implemented a Bayesian optimization approach to efficiently explore the hyperparameter space.
Technical Approach
Key Components:
- Bayesian Optimization: Used Gaussian Process-based optimization to intelligently search the hyperparameter space
- HDBSCAN Clustering: Applied density-based clustering to identify patterns in experimental physics data
- Objective Function: Optimized clustering quality metrics including silhouette score and cluster stability
- Data Processing: Implemented robust preprocessing pipeline using Pandas and NumPy
Optimized Hyperparameters:
min_cluster_size: Minimum number of samples in a clustermin_samples: Number of samples in a neighborhood for a point to be considered core
Results & Impact
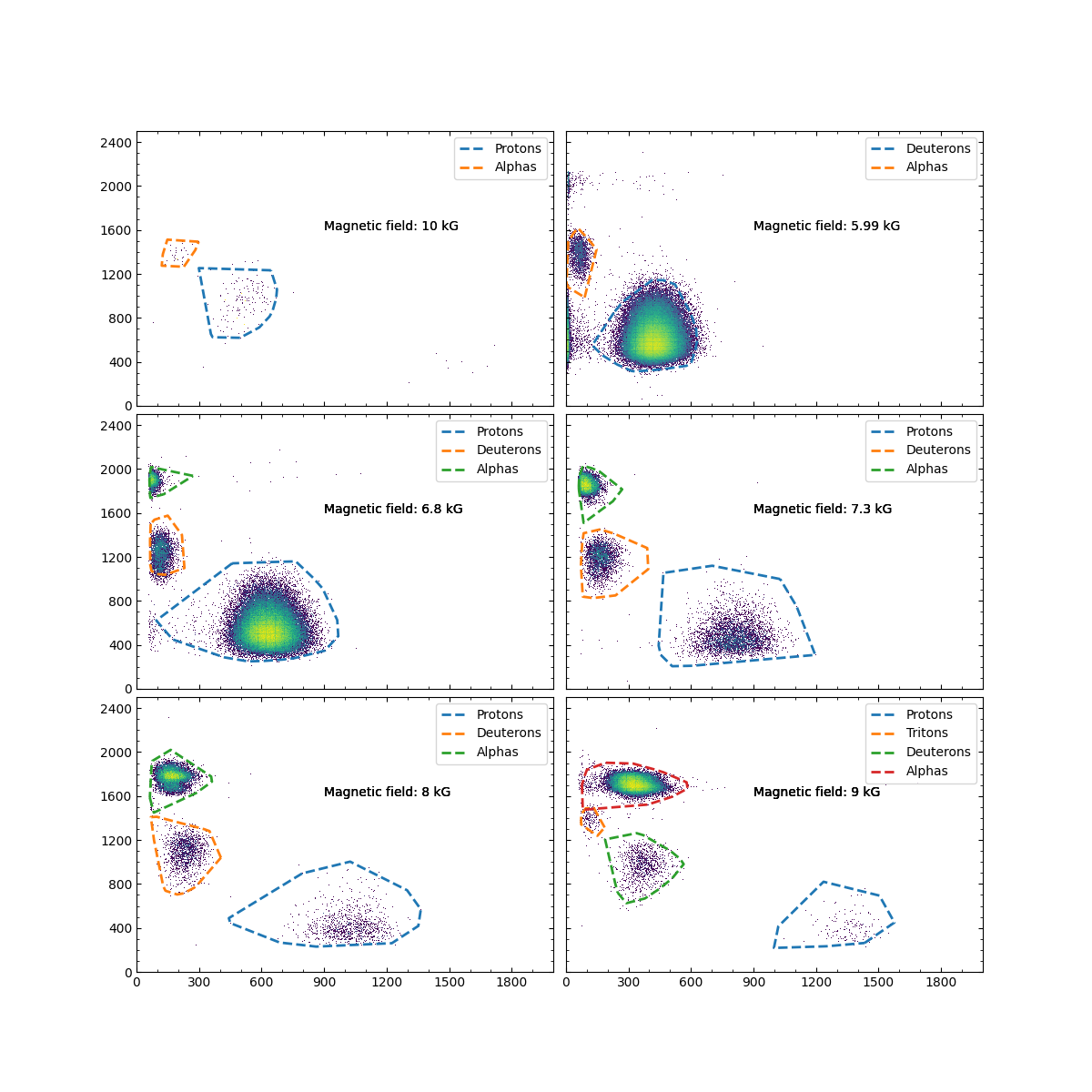
Comparison of clustering results before and after Bayesian optimization
Figure 4 shows the results of the algorithm after running it for each magnetic field setting. Each setting was resampled to 2000 data points except for the 10 KG setting, which had less than 1000 data points in its original set. Overall, the algorithm works very well as it identifies the correct amount of clusters and also prescribes the correct particle label in most field settings. However, proton gates often have a ’tail’ that includes data points to the far left of the dense centroid. This tail may or may not be including noise or particles that are not protons. This could arise from the resampling phase of the algorithm and future analysis will be need to fine tune this and to make sure it does not include incorrect data points. There are misidentifications in the 10 kG and 5.99 kG setting. In the 10 kg setting, the algorithm does not recognize the proton group as it is too sparse. As a result, it incorrectly labels the deuteron group as the proton group and the triton group as the alpha group. In the 5.99 kG setting, the proton group was labeled as the deuteron group and the deuteron group was labeled as the alpha group. It seems the conditions currently set for the algorithm have major limitations when there are only two groups present. Overall, the Bayesian optimization approach significantly improved clustering quality compared to default parameters:
- Reduced optimization time by finding optimal parameters in fewer iterations
- Improved cluster separation and identification of meaningful patterns in SPS experimental data
- Provided a reusable framework for hyperparameter optimization in similar clustering problems